volumegraph.h File Reference
#include <bioprocessing/graph/edgeweightedgraph.h>#include <bioprocessing/graph/setedge.h>#include <bioprocessing/graph/pointvertex.h>#include <aims/connectivity/structuring_element.h>#include <aims/math/mathelem.h>#include <aims/vector/vector.h>#include <cartodata/volume/volume.h>
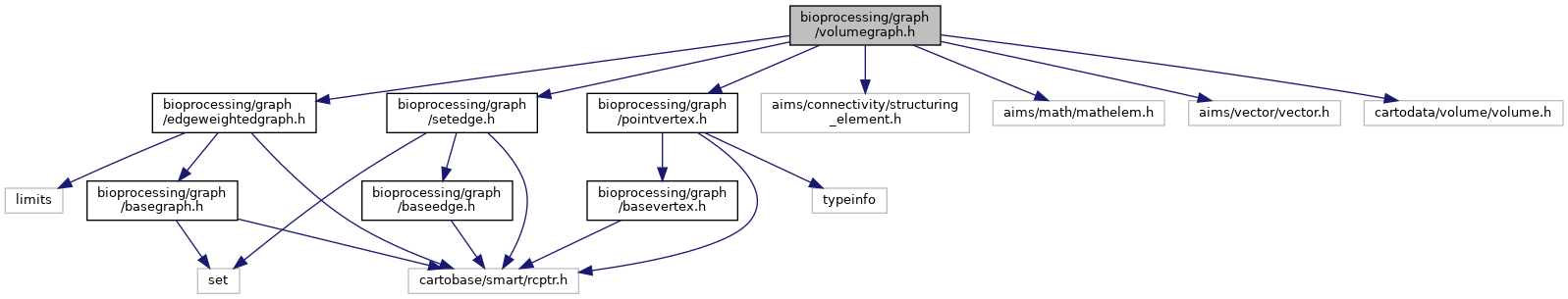
Include dependency graph for volumegraph.h:
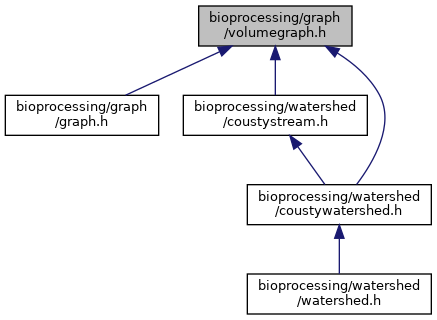
This graph shows which files directly or indirectly include this file:
Go to the source code of this file.
Classes | |
| class | bio::VolumeGraph< T, P > |
| Volume graphs are a graph implementation specifically designed to represent volumes as graphs (vertices are the voxels, and edges are a chosen connectivity). More... | |
| class | bio::VolumeGraphRef< T, P > |
| Reference to a VolumeGraph. More... | |
| class | bio::VolumeGraph< T, P > |
| Volume graphs are a graph implementation specifically designed to represent volumes as graphs (vertices are the voxels, and edges are a chosen connectivity). More... | |
| class | bio::VolumeGraph< T, P >::vertex_const_iterator |
| Iterator on the vertices of a volume graph. More... | |
| class | bio::VolumeGraph< T, P >::vertex_iterator |
| class | bio::VolumeGraph< T, P >::edge_const_iterator |
| Iterator on the edges of a volume graph. More... | |
| class | bio::VolumeGraph< T, P >::edge_iterator |
| class | bio::VolumeGraphRef< T, P > |
| Reference to a VolumeGraph. More... | |
Namespaces | |
| bio | |
