Other tools in the BrainVISA framework¶
HighRes cortex: lamintion of the cortex¶

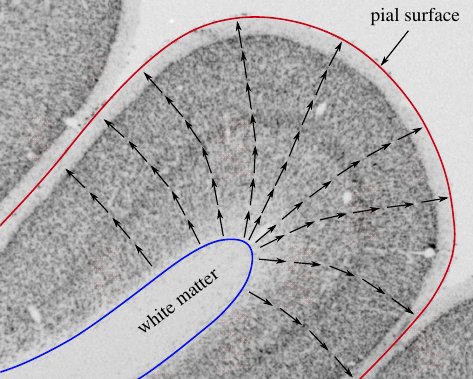
Analysis of the laminar structure of the cortex in high resolution MRI
It is a BrainVisa toolboix, ans also a development library and tools, available on a public sources repository:
https://github.com/neurospin/highres-cortex
See also the development docs.
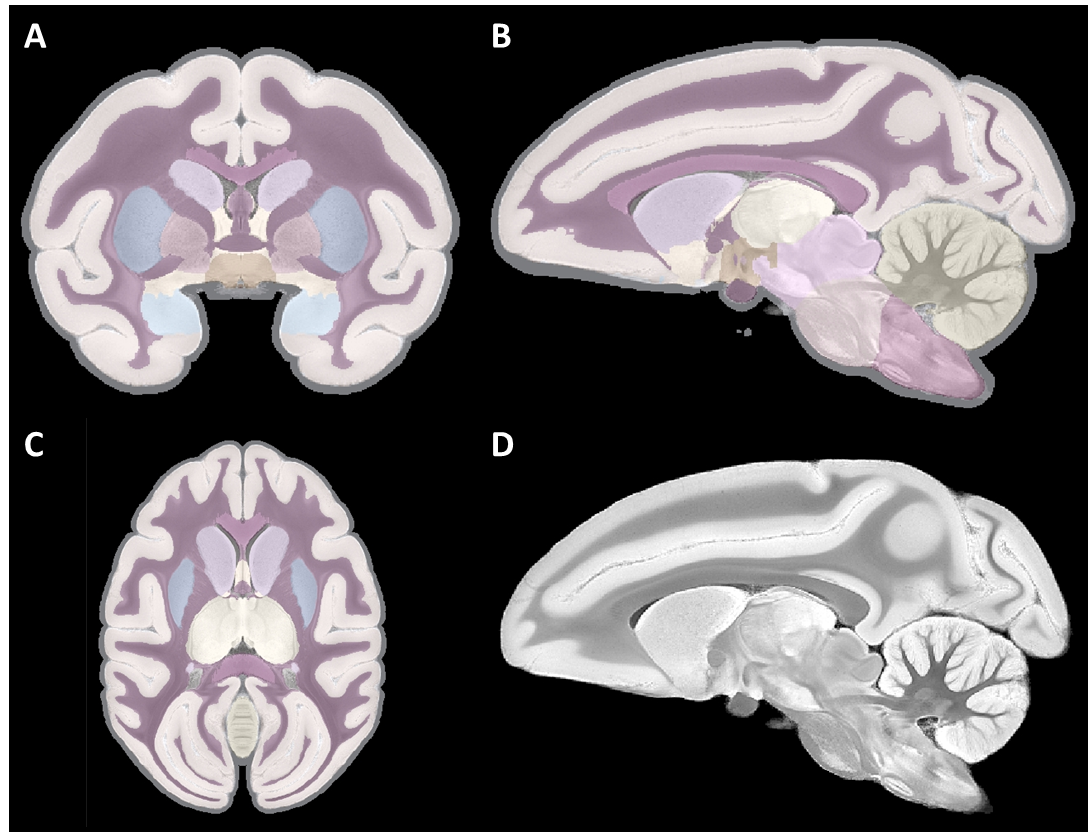
Primate segmentation toolbox¶
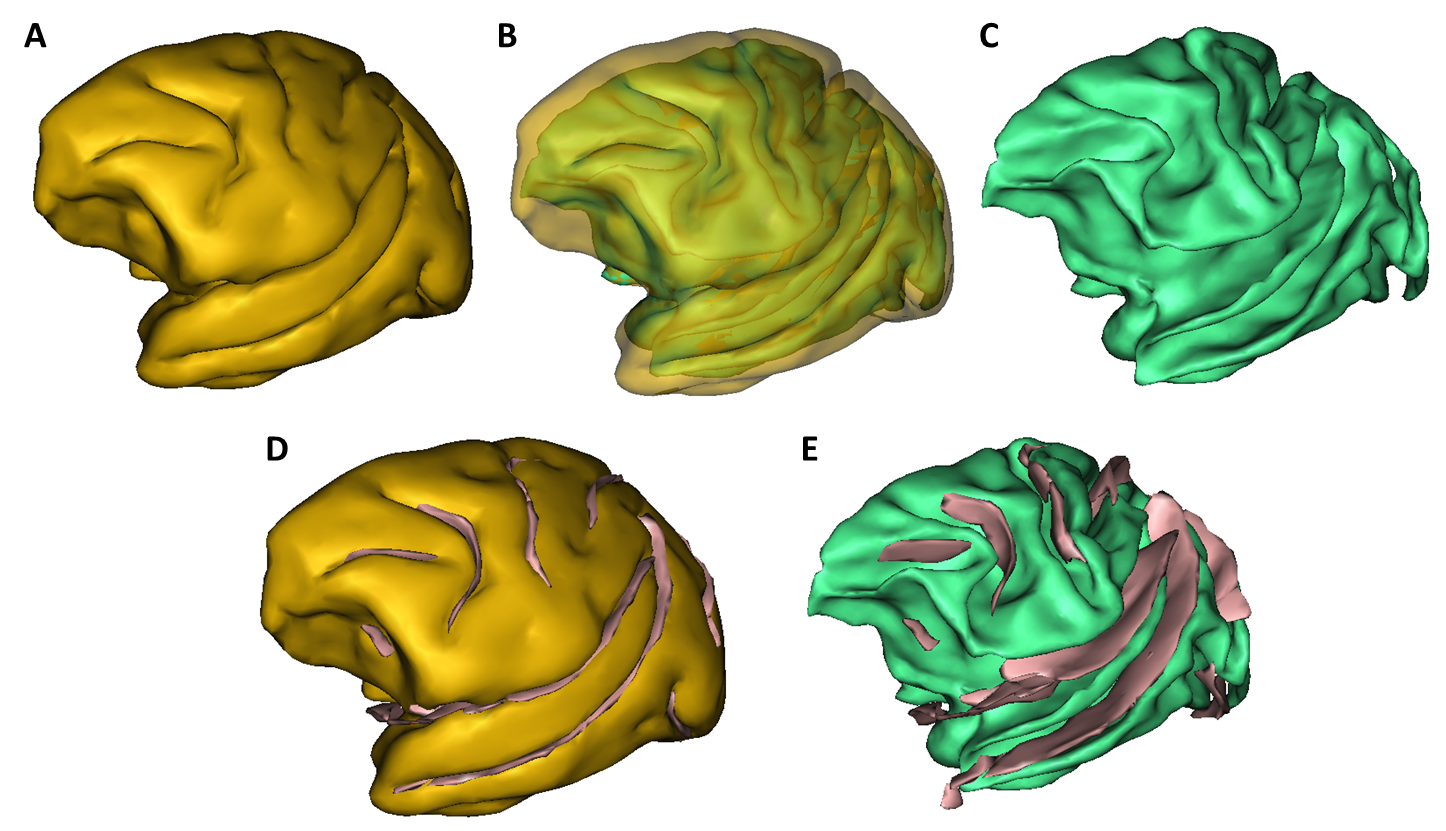
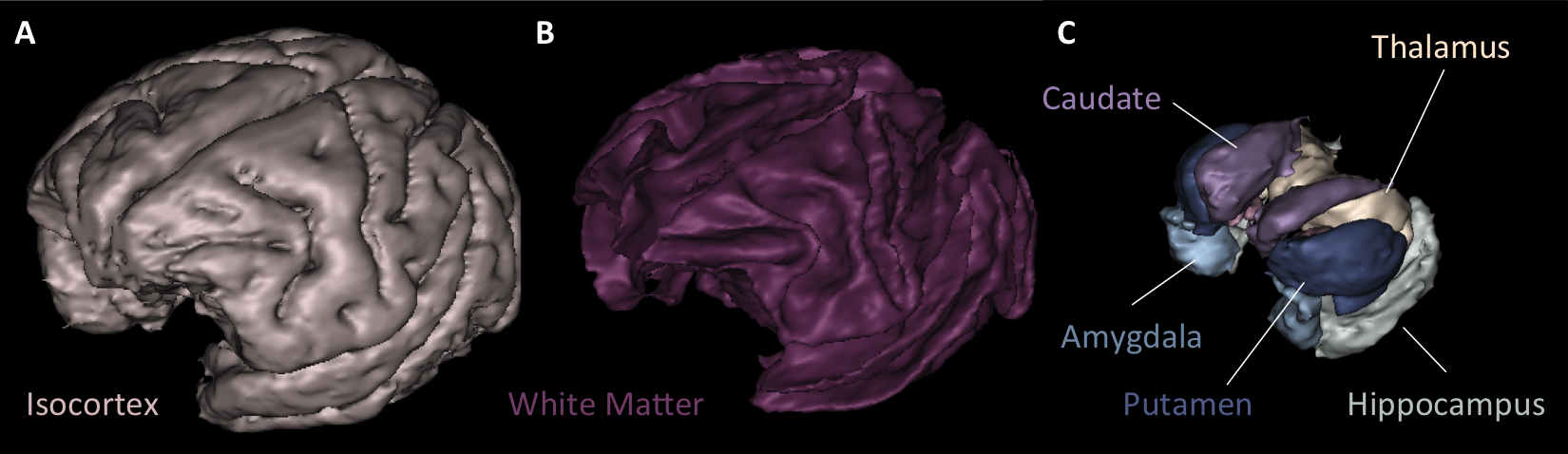
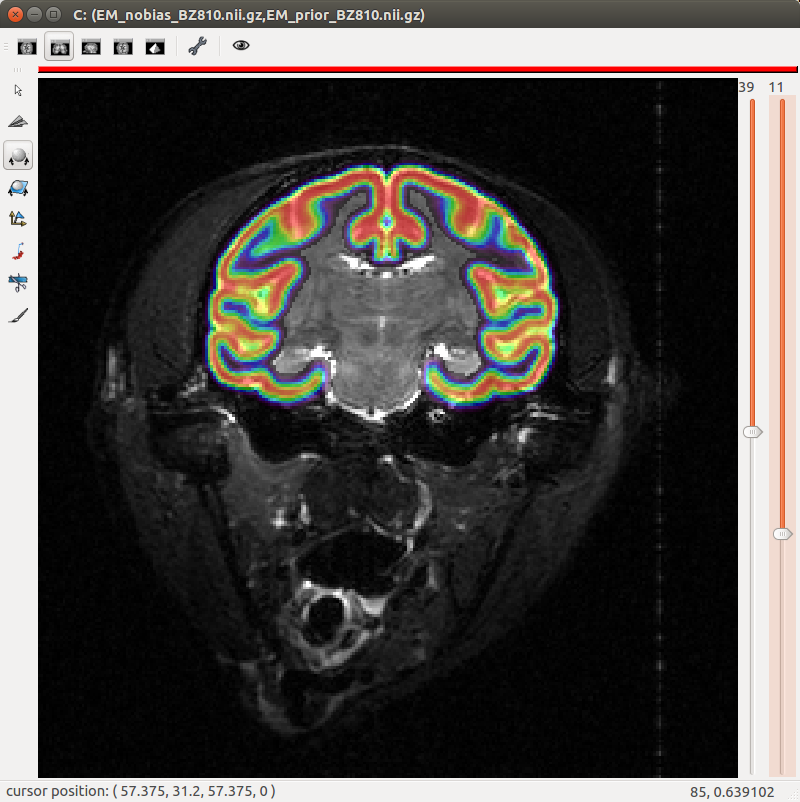
A segmentation toolbox dedicated to primate brain images is developed in MirCen, BioPicsel team. It is making use of some steps of the Morphologist pipeline, but is based on a probabilistic framework. It is designed to process macaque brains with T2 contrasts.
DISCO: constraints-based diffeomorphic registration¶

Diffeomorphic structural-based cortical registration: 3D registration using sulcal constraints.
DISCO toolbox brainvisa help pages
DISCO is developed both by the MeCA research group in Marseille,  Ebrains (formerly the
Ebrains (formerly the  Human Brain Project) and the
Human Brain Project) and the  CATI project labs
CATI project labs
Disco has a long-lasting existence but has finally been publicly released only in BrainVisa 6.0 in 2026.
Champollion: deep learning model for cortical folding¶
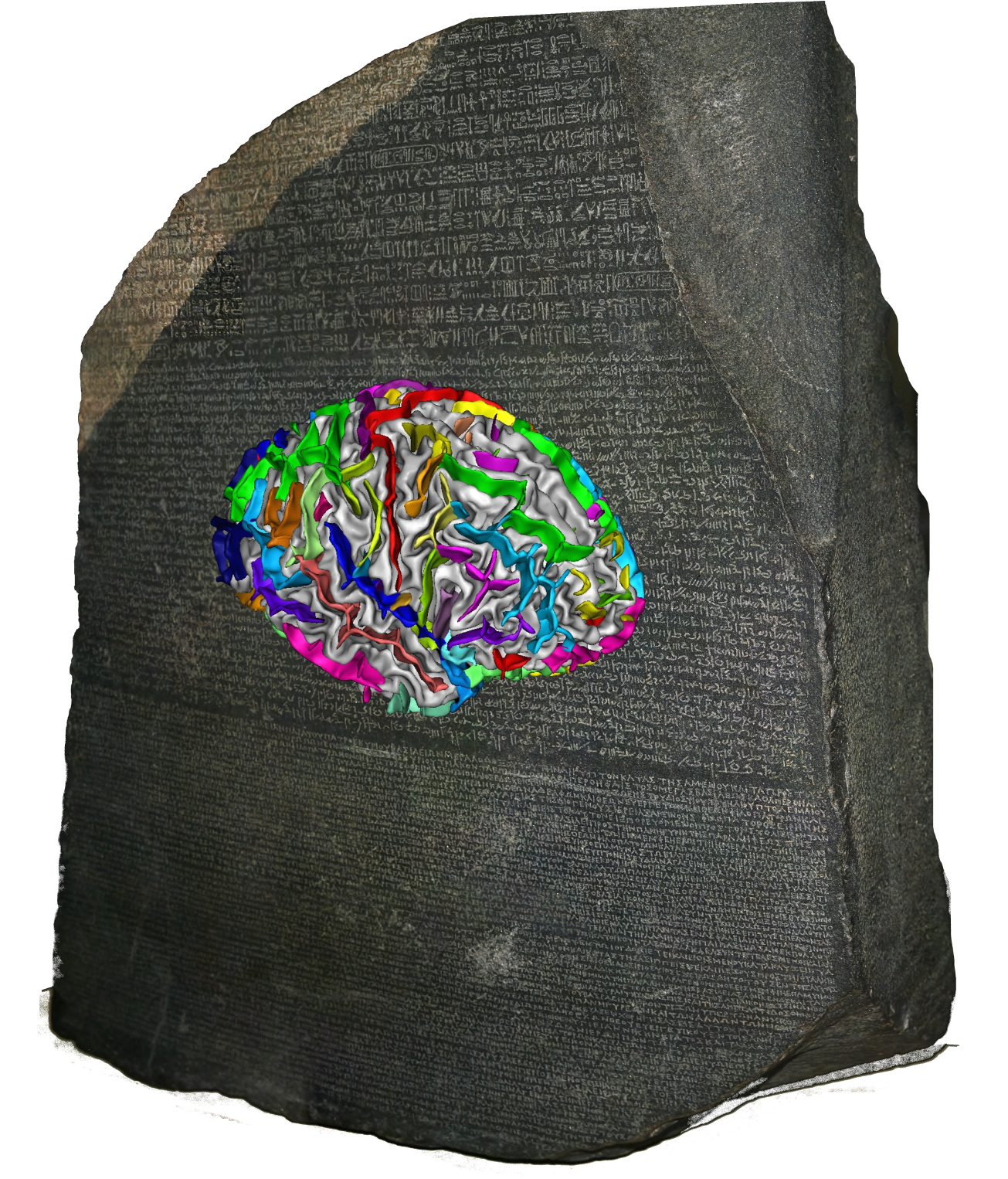

Regional self-supervised learning models of cortical foldiong variability.
New: Released in 2026
Champollion uses self-supervised learning (Barlow Twins) to produce compact vector representations of 28 overlapping sulcal regions per hemisphere (56 total) from 3D brain MRI data. These embeddings capture individual cortical folding variability and can be used for downstream tasks such as classification, clustering, or population-level analysis.
Each model corresponds to one sulcal region and hemisphere. Given a preprocessed brain crop, the model outputs a fixed-size embedding vector per subject.
The model has learned from unlabelled sulcal representations of 40000 brains.
Each region of an indiviudual brain is encoded into a low-dimensional embedding of 32 dimensions per region. Still, this representation contains enough information to reconstruct the original brain folding using a decoder model, and to perform classification or regression tasks for clinical or behavioral variables, using even linear classifiers or regressors when these variables are related to folding.
Champollion takes as input the outputs of the Morphologist toolbox, ie cortical sulci, without labellings.
The complete Champollion pipeline can be found at:
https://github.com/neurospin/champollion_pipeline
Instructions on how to install and use it are found there. More precisely, the Champollion pipeline is made up of 3 sub-pipelines:
Morphologist: MRI segmentation and sulci extraction
Cortical Tiles: split into regions and data preprocessing for Champollion
Champollion: the deep learning model, get embeddings from individual data.
Then the researcher is free to use any statistical toolbox to use the embeddings representations. Examples will be provided on the Champollion Pipelines site.
The pipeline and models can also be found on HuggingFace, with a live demo: