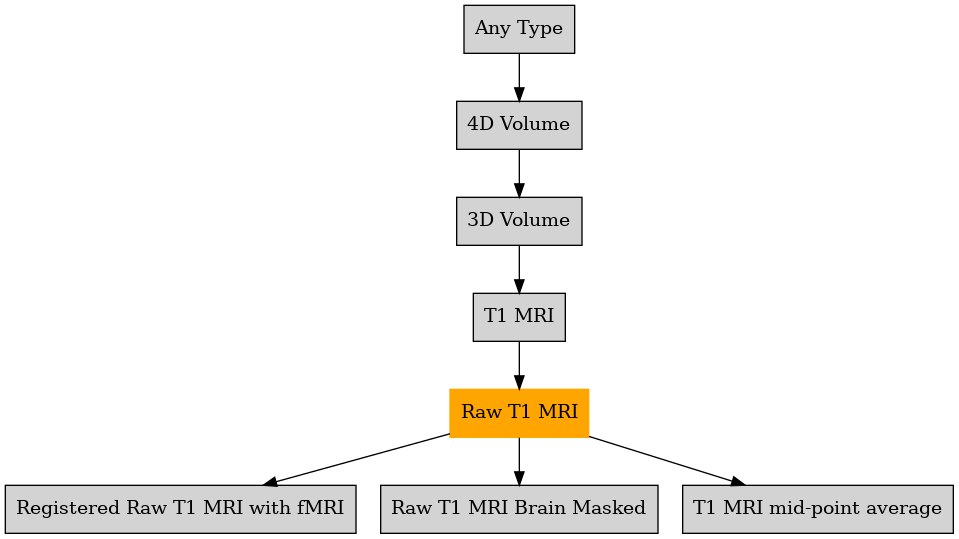

Defined in file : brainvisa/types/builtin.py
01 Create Freesurfer subject from T1 anatomical image
18 Meshes normalization
AC/PC Or Normalization
Add Border To Image
Add gaussian noise in image background
Ana Brain Mask from T1 MRI
Anatomist Show Bias Correction
Anatomist show Commissure coordinates
Anatomist Show White Ridge on T1 MRI
Anatomy Normalization (using AimsMIRegister)
Anatomy Normalization (using Baladin)
Anatomy Normalization (using FSL)
Anatomy Normalization (using FSL) with Re-initialization
Anatomy Normalization (using SPM)
Anatomy Normalization (using SPM) with Re-initialization
Anatomy Normalization (using SPM12)
Anatomy Normalization (using SPM12) with Re-initialization
Anatomy Normalization (using SPM8)
Anatomy Normalization (using SPM8) with Re-initialization
Baladin Normalization Pipeline
CAT12 - Segment
Correction Brain Mask from T1 MRI
Freesurfer / BrainVisa full pipeline
FSL Normalization Pipeline
Import Baby T2 MRI
Import Brain Mask
Import Dicom T1 MRI
Import FreeSurfer grey/white segmentation to Morphologist
Import MNI CIVET Segmentation
Import T1 MRI
Import T1 MRI, longitudinal version
Label volume editor
Localize Talairach Coordinate
Morphologist 2011
Morphologist 2012
Morphologist 2013
Morphologist 2015
Morphologist 2021
Normalization pipeline
Pipeline Baby
Pipeline Baby (Utrecht)
Prepare Subject for Anatomical Pipeline
Read and complete AC-PC files
Reorient Anatomy
Segmentation of Cerebellum, Central Grey Nuclei and Ventricules (Utrecht)
Simplified Morphologist 2015
Skull-stripped brain normalization
Skull-stripping
SPM Normalization Pipeline
spm12 - Segment
spm8 - New Segment
spm8 - VBM Segmentation
Sulcal Lines Extraction Left
Sulcal Lines Extraction Right
T1 Bias Correction
T1 Bias Correction
T1 mapping maps: build using VFA method
T1 Pipeline 2007
Talairach Transformation From Normalization
Transform a commissures AC-PC coordinates
Validation Pipeline
Validation_1 Bias Correction from T1 MRI
View in hippocampic referential
Aperio svs
BMP image
BrainVISA volume formats
DICOM image
Directory
ECAT i image
ECAT v image
FDF image
FreesurferMGH
FreesurferMGZ
GIF image
GIS image
gz compressed ECAT i image
gz compressed ECAT v image
gz compressed GIS image
gz compressed MINC image
gz compressed NIFTI-1 image
gz compressed SPM image
gz compressed VIDA image
Hamamatsu ndpi
Hamamatsu vms
Hamamatsu vmu
JPEG image
Leica scn
MINC image
NIFTI-1 image
PBM image
PGM image
PNG image
PPM image
Sakura svslide
SPM image
TIFF image
TIFF(.tif) image
Ventana bif
VIDA image
XBM image
XPM image
Z compressed ECAT i image
Z compressed ECAT v image
Z compressed GIS image
Z compressed SPM image
Z compressed VIDA image
Zeiss czi
{protocol}/{subject}/anatomy/<subject>
{protocol}/{subject}/anatomy/w<subject>
{protocol}/{subject}/anatomy/<subject>_*
{protocol}/{subject}/anatomy/n<subject>_*
{protocol}/{subject}/anatomy/{acquisition}/<subject>
{protocol}/{subject}/anatomy/{acquisition}/w<subject>
{protocol}/{subject}/anatomy/{acquisition}/<subject>_*
{protocol}/{subject}/anatomy/{acquisition}/n<subject>_*
{protocol}/{subject}/t1mri/{acquisition}/<subject>
{protocol}/{subject}/t1mri/{acquisition}/normalized_{normalization}_<subject>
{center}/{subject}/t1mri/{acquisition}/<subject>
{center}/{subject}/t1mri/{acquisition}/normalized_{normalization}_<subject>
<filename>_t1
normalized_<filename>
sub-{subject}/ses-{session}/anat/t1mri/{acquisition}/<subject>
sub-{subject}/ses-{session}/anat/t1mri/{acquisition}/normalized_{normalization}_<subject>
sub-{subject}/ses-{session}/anat/toto
sub-{subject}/ses-{session}/anat/sub-<subject>_ses-<session>_acq-{acquisition}_T1w
protocol , subject , acquisition , spm_normalized , filename_variable
protocol , subject , acquisition , normalization
time_duration , acquisition_date , time_point , rescan , center , subject , acquisition , analysis , normalization , acquisition_sequence , processing
time_duration , acquisition_date , time_point , rescan , subject , session , acquisition , analysis , normalization
subject , session , acquisition