


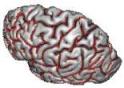
Trigger all segmentation tools for T1-weighted data
This pipeline is now named "Morphologist", and is the main entry point of the "Morphologist" toolbox dedicated to the processing of T1 MRI data. It is the evolution of the older "T1 Pipeline".
Morphologist 2021 does not change in terms of segmentation, but now switches to a CNN-based (Convolutional Neural Networks) sulci recognition method. This step is also now enabled by default.
Older versions of the pipeline are still available in the sub-directory "segmentation Pipeline / Older pipelines", and should normally work as they used to.
News in 2021 vs. 2015:
The sulci recognition sub-pipeline has been enriched with a 3rd generation of methods, based on deep learning CNN models. It performs slightly better than the former SPAM model, and is also way faster. However it needs a fair amount of memory to run (about 6 Gb), or a GPU with Cuda enabled.This pipeline allows to compute:
- either all of what can be done ;
- or a subset oriented towards a specific application :
- Spherical meshes of the cortex (inflation, Cortical Surface Mapping, "primal sketch" of mean curvature, MEG/EEG inverse problem...) ;
- Hemisphere meshes (visualisation, mapping of activations...) ;
- Cortical fold graphs (structural morphometry, constraints for spatial normalisation, surgical planing...).
Before running a pipeline, you have to, either enable a normalization process (needing either SPM or FSL software), or fill in image orientation parameters in the first step, or independently use the Prepare Subject for Anatomical Pipeline process. Be careful that the default recognition method in BrainVISA 4.2 has changed to SPAM recognition with global+local registration, if the corresponding models have been installed.
A pipeline normally deals with both hemispheres, but can be asked to process only one. Processing steps are executed in the following order (except when some results have already been validated/locked or unchecked in the graphical interface) :
- Ensure image orientation and reorient it if needed (Prepare Subject for Anatomical Pipeline) ;
- Computation of a brain mask (Brain Mask Segmentation) ;
- Computation of a mask for each hemisphere (Split Brain Mask) ;
- A grey/white classification of each hemisphere to perform "Voxel Based Morphometry" (Grey White Classification) and spherical triangulation of cortical hemispheres (Grey White Surface) ;
- Spherical triangulation of the external interface of the cortex of one or two hemispheres (Get Spherical Hemi Surface) ;
- Computation of a graph representing the cortical fold topography (Cortical Fold Graph) ;
- Possibly, automatic identification of the cortical sulci (Automatic Sulci Recognition), located in the "sulci" toolbox.
Some of these steps may allow to choose between an older version of the algorithm and a new one. The default settings correspond to the alternative we consider to be the most robust or the most performant, but a different choice may achieve better results on some images.
With some practice, you may play iteratively with the pipelines. For instance, if you have trigerred a "sub-pipeline" only, you may ask for more without having to recompute the whole stuff. For that purpose, you can lock the first results using the files locking system (right click on the data parameter name and select "lock" in the menu), or alternatively use the old BrainVISA validation tools.It is to be noted that since version 2012, the pipeline has been made robust enough to run all steps, with the default settings and without manual intervention, on a large majority of cases.
The different active pipelines are the following (you may find other ones elsewhere when the input is not the raw T1-weighted MR scan):





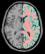

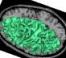
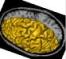
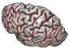
Efficient use of the pipeline:
If you want to process a large population of brains, we advise the following strategy :
- Select a few representative brains, and process them with the default parameters. If the results are not so good, several levels of tuning can be tried in order to achieve acceptable results :
- You can change the field rigidity parameter in the Bias Correction. If the contrast is bad, you can increase this parameter and vice-versa.
- In the Brain Mask Segmentation, you can increase the erosion size.
- If your image's contrast is bad or if your image is noised, it can be better to reduce the bary factor parameter in the Split Brain Mask, which will reduce the threshold between the grey matter and white matter.
If you are not unlucky, you should find a good solution with this, otherwise it may be interesting to derive your own pipeline using low-level bricks, Vip library and maybe your own local tools. This is what is done for example in Marseille fMRI site because of the 3T large bias field: http://marspack.free.fr/.
- After the tuning stage, we suggest to perform the processing in two stages :
- From raw MR data to the brain mask : It can be iterated on all your brains. Then check the results one by one and if still some data's results are not acceptable try some of the alternatives above. (For the very recalcitrant brains, you could perform some manual corrections with Anatomist drawing's capacities : Label volume editor.)
- Any further processing (meshes, graphs, etc.) : You have first to lock the results of the previous stages, for them not to be superseded by the general pipeline treatment. Use : Ana Validate All. Then you can iterate the pipeline you like and go to sleep.
Visual control at databases scale
After running the Morphologist pipeline on a set of subjects, it is highly recommended to visually inspect the results: a pipeline which runs to the end does not imply that it has produced good results...
A new visual control module has been released since BrainVisa / Morphologist version 4.3: SnapBase. This tool allows to perform snapshots of the visualization of some data, under given points of view, on entire databases, so as to present the results for all subjects together. This allows to spot, a posteriori, very quickly and easily, some subjects for which the analysis has largely failed, amongst large amounts of subjects.












t1mri: Raw T1 MRI ( input )
perform_normalization: Boolean ( input )
anterior_commissure: Point3D ( optional, input )
posterior_commissure: Point3D ( optional, input )
interhemispheric_point: Point3D ( optional, input )
left_hemisphere_point: Point3D ( optional, input )
t1mri_nobias: T1 MRI Bias Corrected ( output )
histo_analysis: Histo Analysis ( output )
split_brain: Split Brain Mask ( output )
left_graph: Cortical folds graph ( output )
right_graph: Cortical folds graph ( output )
perform_sulci_recognition: Boolean ( input )
left_labelled_graph: Labelled Cortical folds graph ( optional, output )
right_labelled_graph: Labelled Cortical folds graph ( optional, output )
Toolbox : Morphologist
User level : 0
Identifier :
morphologistFile name :
brainvisa/toolboxes/morphologist/processes/morphologist.pySupported file formats :
t1mri :gz compressed NIFTI-1 image, Aperio svs, BMP image, DICOM image, Directory, ECAT i image, ECAT v image, FDF image, FreesurferMGH, FreesurferMGZ, GIF image, GIS image, Hamamatsu ndpi, Hamamatsu vms, Hamamatsu vmu, JPEG image, Leica scn, MINC image, NIFTI-1 image, PBM image, PGM image, PNG image, PPM image, SPM image, Sakura svslide, TIFF image, TIFF image, TIFF(.tif) image, TIFF(.tif) image, VIDA image, Ventana bif, XBM image, XPM image, Zeiss czi, gz compressed MINC image, gz compressed NIFTI-1 imaget1mri_nobias :gz compressed NIFTI-1 image, BMP image, DICOM image, Directory, ECAT i image, ECAT v image, FDF image, GIF image, GIS image, JPEG image, MINC image, NIFTI-1 image, PBM image, PGM image, PNG image, PPM image, SPM image, TIFF image, TIFF(.tif) image, VIDA image, XBM image, XPM image, gz compressed MINC image, gz compressed NIFTI-1 imagehisto_analysis :Histo Analysis, Histo Analysissplit_brain :gz compressed NIFTI-1 image, BMP image, DICOM image, Directory, ECAT i image, ECAT v image, FDF image, GIF image, GIS image, JPEG image, MINC image, NIFTI-1 image, PBM image, PGM image, PNG image, PPM image, SPM image, TIFF image, TIFF(.tif) image, VIDA image, XBM image, XPM image, gz compressed MINC image, gz compressed NIFTI-1 imageleft_graph :Graph and data, Graph and dataright_graph :Graph and data, Graph and dataleft_labelled_graph :Graph and data, Graph and dataright_labelled_graph :Graph and data, Graph and data