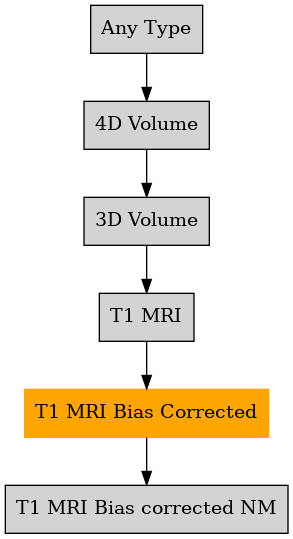

Defined in file : brainvisa/types/builtin.py
2D Geodesic Primal Sketch
Ana Brain Mask from T1 MRI
Ana Compute Hemi Grey White Classification
Ana Get Opened Hemi Surface
Ana Get Opened Whole Brain Surface
Ana Get Spherical Cortical Surface
Ana Split Brain from Brain Mask
Anatomist Show Bias Correction
Anatomist Show Brain Mask on T1 MRI
Anatomist Show Fold Graph
Anatomist Show Grey White on T1 MRI
Anatomist Show GreyWhite Snake on T1 MRI
Anatomist Show Gyrus Graph
Anatomist Show Hemisphere With MRI
Anatomist Show Nucleus Graph
Anatomist Show Split Brain on T1 MRI
Anatomist Show White Matter With MRI
Brain Detection
Brain Detection (Utrecht)
Brain Mask Segmentation
CAT12 - Segment
Change Template Referential
Combine Bundles
Compute Brain Mask
Compute Cortex image
Correction Brain Mask from T1 MRI
Correction Split Brain from Brain Mask
Cortical Fold Arg (3.0)
Cortical Fold Graph (3.1+)
Cortical Fold Graph (general)
Cortical Fold Graph (one hemisphere)
Cortical Fold Graph 2007-2011 (general)
Create Cortical Constraints Texture
Create DISCO deformation field from subjectA to subjectB
Create DISCO+DARTEL deformation field from subjectA to subjectB
Create histo analysis manually
Create Left Hemisphere Cortical Constraints Texture
Create Right Hemisphere Cortical Constraints Texture
Create Sulcus Label Volume
Edit histo analysis
Get Hemi Surface
Get Spherical Hemi Surface
Graph Structure (3.0)
Graph Structure (3.1)
Grey White Classification 2012
Grey white Interface
Grey White Surface 2012
Gyral Parcellation
Head Mesh
Hemisphere Grey White Classification
Hemisphere Grey White Topological Correction
Hemisphere Pial Mesh
Hemisphere Sulci Skeleton
Histogram analysis
Histogram analysis
Histogram Analysis
Import FreeSurfer grey/white segmentation to Morphologist
Import MNI CIVET Segmentation
Import Vois
Inner Surface Initialization
Inner Surface Initialization (Utrecht)
Insular Pole Projection
Intensity normalisation PET Pipeline (spm12)
Intensity normalisation PET Pipeline (spm8)
Manual Drawing of Cerebellum and Segmentation of Central Grey Nuclei from 2 Orthogonal Slices
Mesh deformation component
Morphologist 2011
Morphologist 2012
Morphologist 2013
Morphologist 2015
Morphologist 2021
Normalize intensities of a T1 MRI
Pipeline Baby
Pipeline Baby (Utrecht)
Quality control for Pet with MRI Pipeline
Quality control for SUVSUVR compute stat
Quality control for SUVSUVR MNI space pipeline
Quality control for SUVSUVR native space pipeline
Quality control for SUVSUVR PET parametric
Quality control for SUVSUVR Preprocess
Right Insular Pole Projection
ROI based analysis PET Pipeline (spm12)
ROI based analysis PET Pipeline (spm8)
segmentation choice (spm8)
Simplified Brain Detection (Utrecht)
Simplified Morphologist 2015
Split Brain Mask
Split Brain Mask
spm12 - Segment
spm8 - New Segment
spm8 - VBM Segmentation
Sulcal Lines Extraction
Sulcal Lines Extraction With Options
Sulcal Parcellation
Sulci Voronoi
Sulcus Parameterization
Sulcus Parameterization 2015
Surface Deformation
Surface Deformation (Utrecht)
Swap Bias Corrected MRI and Simulated MRI
T1 Bias Correction
T1 Bias Correction
T1 Pipeline 2007
Transform fibre bundles between subjects
Transform fibre bundles into the common space
Transform graph between subjects
Transform graph into the common space
Transform mesh between subjects
Transform mesh into the common space
Transform volume between subjects
Transform volume into the common space
Validation Pipeline
Validation_1 Bias Correction from T1 MRI
Validation_2 Nobias Histo analysis
Validation_3 Brain Mask from T1 MRI
Validation_4 Split Brain from Brain Mask
Vip Bias Correction
Vip Get Brain
Vip Histogram analysis
Vip Split Brain
Aperio svs
BMP image
BrainVISA volume formats
DICOM image
Directory
ECAT i image
ECAT v image
FDF image
FreesurferMGH
FreesurferMGZ
GIF image
GIS image
gz compressed ECAT i image
gz compressed ECAT v image
gz compressed GIS image
gz compressed MINC image
gz compressed NIFTI-1 image
gz compressed SPM image
gz compressed VIDA image
Hamamatsu ndpi
Hamamatsu vms
Hamamatsu vmu
JPEG image
Leica scn
MINC image
NIFTI-1 image
PBM image
PGM image
PNG image
PPM image
Sakura svslide
SPM image
TIFF image
TIFF(.tif) image
Ventana bif
VIDA image
XBM image
XPM image
Z compressed ECAT i image
Z compressed ECAT v image
Z compressed GIS image
Z compressed SPM image
Z compressed VIDA image
Zeiss czi
./nobias_<filename>
{center}/{subject}/t1mri/{acquisition}/{analysis}/nobias_<subject>
{center}/{subject}/longitudinal_preprocessings/spm12Serial/avg_{acquisition_sequence}/{analysis}/<subject>_{acquisition}_to_avg_<acquisition_sequence>_bias_corrected
{center}/{subject}/longitudinal_preprocessings/spm12Pairwise/avg_{acquisition_sequence}/{analysis}/<subject>_{acquisition}_to_avg_<acquisition_sequence>_bias_corrected
{center}/{subject}/spm/cat12Segment/{acquisition}/{analysis}/mri/m<subject>
{center}/{subject}/spm/cat12Segment/{acquisition}/{analysis}/mri/wm<subject>
{center}/{subject}/spm/spm12Segment/{acquisition}/{analysis}_LDW_from_t1mri_to_{template}/<subject>_bias_corrected
{center}/{subject}/spm/spm12Segment/{acquisition}/{analysis}_LDW_from_t1mri_to_{template}/<subject>_bias_corrected_using_affine_registered
{center}/{subject}/spm/spm12Segment/{acquisition}/{analysis}_LDW_from_t1mri_to_{template}/<subject>_bias_corrected_using_warped_without_modulation
{center}/{subject}/spm/spm12Segment/{acquisition}/{analysis}_HDW_from_t1mri_to_{template}/<subject>_bias_corrected_using_warped_without_modulation
{center}/{subject}/spm/spm8VBMSegmentation/{acquisition}/{analysis}_LDW_from_t1mri_to_{template}/<subject>_bias_corrected
{center}/{subject}/spm/spm8VBMSegmentation/{acquisition}/{analysis}_LDW_from_t1mri_to_{template}/<subject>_bias_corrected_using_affine_registered
{center}/{subject}/spm/spm8VBMSegmentation/{acquisition}/{analysis}_LDW_from_t1mri_to_{template}/<subject>_bias_corrected_using_warped_without_modulation
{center}/{subject}/spm/spm8VBMSegmentation/{acquisition}/{analysis}_HDW_from_t1mri_to_{template}/<subject>_bias_corrected_using_warped_without_modulation
{center}/{subject}/spm/spm8NewSegment/{acquisition}/{analysis}_LDW_from_t1mri_to_{template}/<subject>_bias_corrected
{center}/{subject}/spm/spm8NewSegment/{acquisition}/{analysis}_LDW_from_t1mri_to_{template}/<subject>_bias_corrected_using_affine_registered
{center}/{subject}/spm/spm8NewSegment/{acquisition}/{analysis}_LDW_from_t1mri_to_{template}/<subject>_bias_corrected_using_warped_without_modulation
{center}/{subject}/spm/spm8NewSegment/{acquisition}/{analysis}_HDW_from_t1mri_to_{template}/<subject>_bias_corrected_using_warped_without_modulation
{protocol}/{subject}/anatomy/nobias_<subject>
{protocol}/{subject}/anatomy/nobias_<subject>_*
{protocol}/{subject}/anatomy/nnobias_<subject>_*
{protocol}/{subject}/anatomy/{acquisition}/nobias_<subject>
{protocol}/{subject}/anatomy/{acquisition}/nobias_<subject>_*
{protocol}/{subject}/anatomy/{acquisition}/nnobias_<subject>_*
{center}/{subject}/{analysis}/{acquisition}/using_LDW_from_t1mri_to_{template}/<subject>_bias_corrected
{center}/{subject}/{analysis}/{acquisition}/using_LDW_from_t1mri_to_{template}/<subject>_bias_corrected_affine_registered
{center}/{subject}/{analysis}/{acquisition}/using_LDW_from_t1mri_to_{template}/<subject>_bias_corrected_warped_without_modulation
{center}/{subject}/{analysis}/{acquisition}/using_HDW_from_t1mri_to_{template}/<subject>_bias_corrected_warped_without_modulation
{protocol}/{subject}/t1mri/{acquisition}/{analysis}/nobias_<subject>
{center}/{subject}/{analysis}/{acquisition}/using_LDW_from_t1mri_to_{template}/<subject>_bias_corrected
{center}/{subject}/{analysis}/{acquisition}/using_LDW_from_t1mri_to_{template}/<subject>_bias_corrected_affine_registered
{center}/{subject}/{analysis}/{acquisition}/using_LDW_from_t1mri_to_{template}/<subject>_bias_corrected_warped_without_modulation
{center}/{subject}/{analysis}/{acquisition}/using_HDW_from_t1mri_to_{template}/<subject>_bias_corrected_warped_without_modulation
center , subject , processing , acquisition , analysis , segmentation_method , tracer , template , rescan , acquisition_date , time_point , time_duration , warping_method , acquisition_sequence , transformation , space
center , subject , analysis , acquisition , template , transformation , warping_method , protocol , filename_variable
protocol , subject , acquisition , analysis , center , template , transformation , warping_method